3D Cell culture & Microfluidics
Recently, microfluidics and 3D cell culture technologies have merged, to increase the potential of in vitro modeling, in order to get even closer to Nature. The successful control of the cellular environment thanks to microfluidics in a 3D cell culture environment allows the study at the single-cell scale with mechanical control of cell shape, physicochemical properties, shear-stress, the rigidity of substrates, temperature and gas gradient. The perpetual increase of knowledge in histology, physiology and in technology, allow us to drastically improve our tools to mimic the living organisms.
Why do we need alternative cell culture models ?
Nowadays, 3D cell culture microfluidic devices are used in two main fields: as in vitro models for fundamental research, drug trial, toxicity assays… or in cell therapy to manufacture tissue prior to implantation (Fig 1). For instance regenerative medicine, which is still in the early stage of development, consists in repairing lesions or diseased organs by replacing the damaged parts with new tissue grown in the lab. This approach follows the principle of cell therapy, and uses stem cells, undifferentiated or de-differentiated cells cultured in vitro, to then transplant them in vivo. The development of these technologies will have a huge impact on healthcare as a solution for many currently untreatable diseases : heart failure, kidney failure, genetic disorders, orphan diseases, cancer…
Cell Therapy
In vitro Research Model
Regenerative medicine
Stem Cell Culture
Drug Discovery
Cancer Study
BioTech & Pharma Indutry
Research Labs & Institutes
Hospital & Diagnostic labs
Fig 1: 3D cell culture fields of applications and players
How can 3D cell culture improve the in vitro model ?
In vivo, cells are surrounded by a well-designed extracellular matrix (ECM) structure. The ECM is composed of macromolecules secreted by the cells (e.g., collagen, elastin, glycosaminoglycan, glycoprotein). Its composition is adapted to each cell type and cell functions, in terms of elasticity, porosity, and roughness rigidity. The ECM is very abundant in conjunctive tissue, but weaker between epithelial cells. Traditional 2D in vitro cell culture models, consisting in growing monolayer of cells on a flat surface, do not allow the integration of ECM. Moreover, cells grown in 2D present different morphology, gene and protein expression, and response to stimuli as compared to 3D in vitro or in vivo culture. These results indicate that the 2D monolayer is not well suited to perform relevant biological assays.
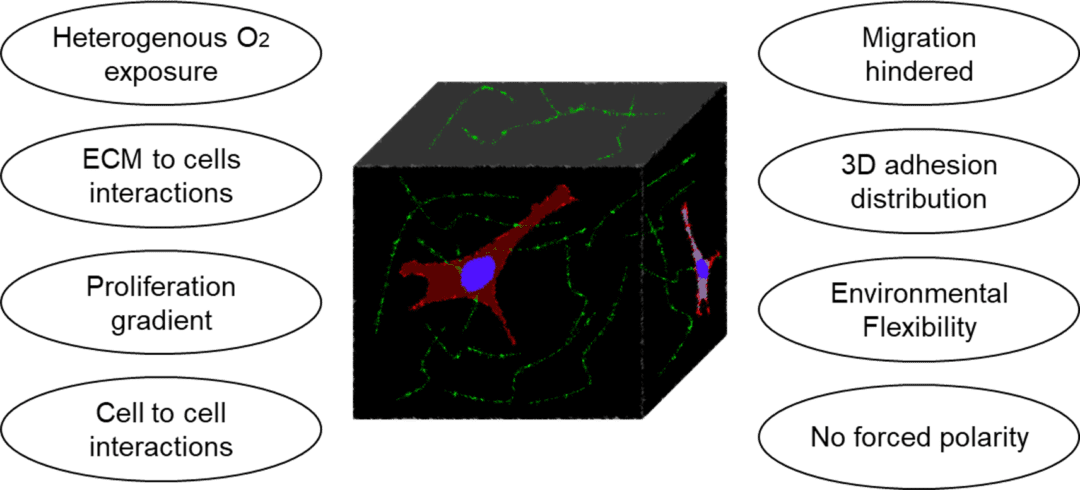
The challenges of novel cell culture techniques is therefore to develop a better suited in vitro cell culture method recapitulating the aforementioned characteristics of the cells and associated ECM. These models also need to be adapted to each organ or biological phenomenon investigated.
In 2019, a Team from the Department of Chemistry in Saint Louis University, USA published an article reviewing the current practices to incorporate 3D culture in microfluidic design. They compared many different approach and classified them into four different methods (Fig 3), presented here [1].
Four 3D cell culture methods
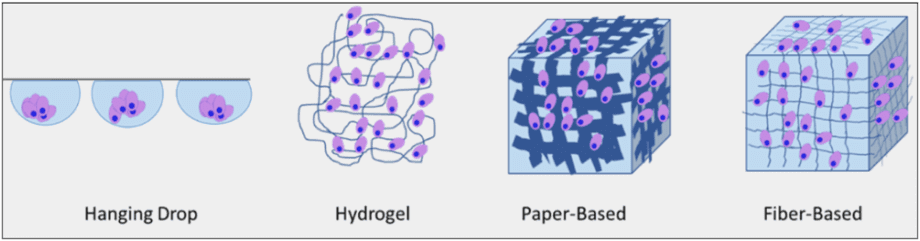
Fig 3: Four types of 3D cell culture method: Hanging Drops, cells are floating in a medium droplet. Hydrogels are made of polymerized hydrophilic polymers. Paper-based scaffolds give cells a 3D cellulose architecture. Fiber-based scaffolds are made of nano-fiber of different sizes. i
The Hanging Drop
This method is a sub-type of suspension cell culture, and is only suited for non-adherent cell lines. Droplets with fine-tuned sizes are suspended on a scaffold. In each droplet, single-cells aggregate into a homogeneous colony. The use of microfluidics allows here to generate a high throughput of aggregates, as well as a continuous perfusion of fresh media. In 2016, Huei-Wen Wu and his team from Taiwan built a device to pump solution through the droplets thanks to a pressure difference between two reservoirs, eliminating the need for an external pumping system (e.g., syringe pump) (Fig. 4). In this instance, they used the device to grow artificial mouse embryoid bodies (EBs) from 3D clusters of pluripotent stem cells. [2]
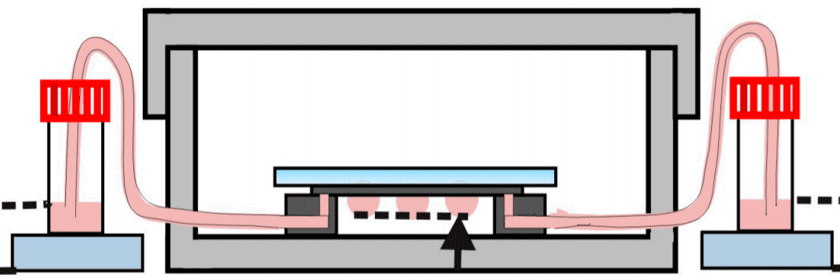
Fig 4: Hanging drop device: EBs-on-a-chip(ii) Schematic representation of the microfluidic device of hanging drops connected to the reservoirs which allow the renewal of the culture medium by pressure difference.
Wu et al., build a polydimethylsiloxane (PDMS) microfluidic chip from SU-8 mold on a glass surface. They optimized pressure conditions to allow a medium exchange of the hanging drops. In the end, they manage to grow multiple embryoid bodies and perform immuno-staining to characterize them.
Hydrogels
Hydrogel solutions are hydrophilic networks of polymers and macromolecules of different composition, which under the right condition, can crosslink into a porous gel, with properties similar to one of the extra cellular matrix. These hydrogels are easy to produce and are often used as alternative to ECM to grow cells in a 3D configuration. Cells will grow and migrate inside the gel, and will be surrounded by small molecules increasing the surface exchanges compare to 2D cell culture. Hydrogels can be made of synthetic (Polyacrylamide and Polyethylene glycol) or natural polymers from the ECM (collagen, fibrin, hyaluronic acid). The composition of hydrogel is adapted to the cell types studied, by tuning the substrate thickness, porosity, and overall stability.
The main drawbacks of hydrogels are their lack of stability, in particular when prepared from natural polymers. Although, this technique did not provide a rigid scaffold that can be useful for certain tissue modeling.
Paper-based Scaffold
Paper is a porous, biocompatible, stackable and foldable material onto which cells can adhere and grow. Paper sheets have been employed to generate complex 3D structures, and mimic the cell environments. Paper-based scaffolds are inexpensive and commercially available. The cellulose network provides a rigid architecture for cells to grow. These materials are very stable and can undergo several surface treatments: chemical treatment, plasma, or sterilization. However, the cellulose matrix mimics poorly the ECM composition. The fiber is much bigger: >1µm compared to 500nm in nature. To include a paper based structure into a microfluidic platform, the current most common approach is hydrophobic wax printing, and these devices are usually coupled with hydrogels.
Tao et al build a paper-based platform where cells are seeding directly into the cell culture areas connected by two channels to a medium reservoir[3].
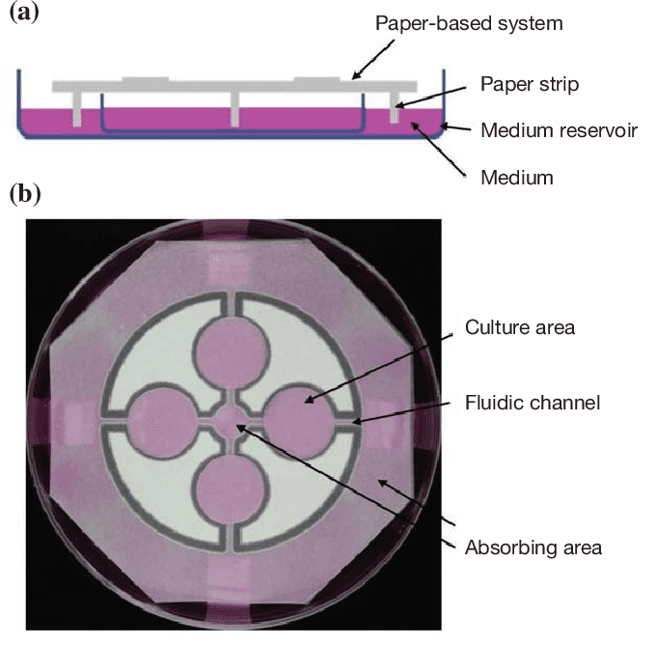
Fig 5: Paper-based cell culture system. (A)The paper sheet is suspending above a medium reservoir with paper strips connecting the two. Fresh medium is passively circulating from the reservoir to the culture areas through the strip. (B)Four culture areas were designed in a T-organization by wax printing. 2 fluidic channels are connecting between each culture area and 2 absorbing areas.[3]
In this microfluidic cellulose filter paper device, medium and other substances can be passively perfused to the culture areas. The team manage to perform analyses of cyto-compatibility, cell proliferation, morphology, assess the potential of the paper-based system. This system can be used for a wide range of assays with applications in drug screening and cell biology.
“This work demonstrated a paper-based cell culture microfluidic system, and the system is inexpensive, disposable, and compatible to the existing culture facility.”
Tao et al., 2015
Fiber scaffold
This technology combines the advantages of hydrogels and paper-based cell culture devices. Fibers that compose the scaffold are produced by several techniques: melt, blow, and electrospinning. Fibers can be natural or synthetic, from different sizes (10µM to nm) mimicking the extracellular matrix. All parameters can be tunned at will and favor stability. Currently, most 3D cell culture devices incorporating fiber scaffolds are made of PDMS. Lee et al. studied human mesenchymal stem cells. They manage to increase their proliferation using acrylic acid modified polyurethane nanofiber in a PDMS chip.[4]
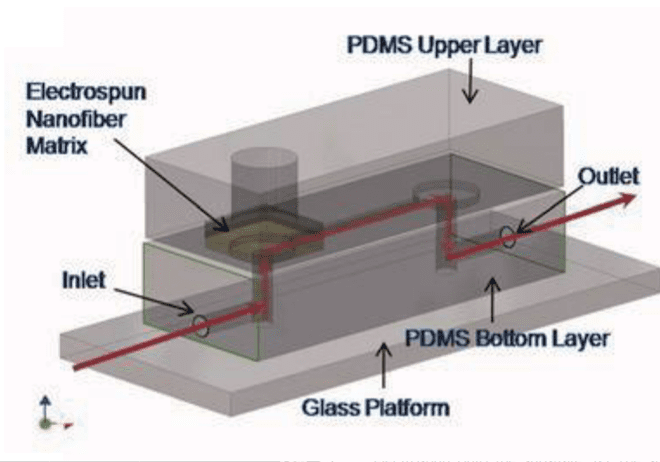
This team has developed a microfluidic platform that has the ability to control biochemical and biomechanical forces in a three-dimensional scaffolding coupled with image acquisition. They were able to assess and quantify capillary growth and endothelial cell migration from a cell monolayer. The response of endothelial cells placed in co-culture (cancer or smooth muscle cells) was studied. The results presented show the capabilities of this technology to study cell morphogenesis while having the advantage of improved imaging and internal biological controls.
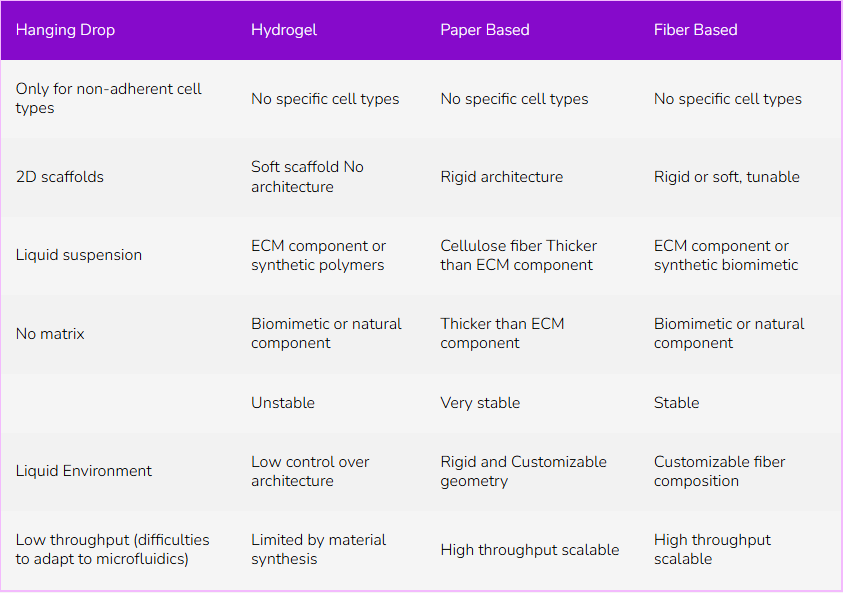
Table 1: Comparison of 3D cell culture technics
Microfabrication challenges
The different types of fibers used in the 3D device have their characteristics, pros, and cons. Various parameters can be adjusted to better suit the needs of the cell type being studied. However, this development can be long, complicated, and requires an extensive knowledge of the tissues and the cell studied. Moreover, most investigations involving microfluidics platforms currently use PDMS as a construction material. This material, although relatively easy to handle on the small scale, with a straightforward protocol, is relatively expensive compared to plastics and other polymers.
For the device replication, PDMS devices are prepared using a soft lithography protocol, by which the polymers are poured on silicon and SU-8 wafer molds, containing a negative of the desired structure. These molds are fragile, expensive and require a long process to be built. Additionally the soft lithography approach does not allow for a high reproducibility of the device. All theses challenges prevent the PDMS from being used at the industrial scale.
Finally, the last step of PDMS microfluidic device preparation consists in sealing the channels and closing the system. This step requires an expensive plasma cleaner, and relies and the plasma activation of PDMS to seal the device. This approach has shown uneven levels of efficacy, leading to leaky devices. Moreover, the plasma activation can induce changes the functionalization of the microchannels walls, affecting the ensuing experiments.
At Eden Tech we focus our work to improve the daily experience in terms of material and facilities for Biology Lab: Transfer microfluidic out of the cleanroom. We propose safer materiel that requires less solvent, easier manipulation…
FlexdymTM is a Gaz permeable, biocompatible, and transparent thermoplastic and is proposed as a cheap alternative to PDMS. Microfabrication is easily performed by hot embossing. FlexdymTM devices are replicated in few minutes by the Sublym hot embossing device. No toxic solvents or particular heavy infrastructures are required. Finally, EpoxymTM helps you reproduce with high fidelity your silicon wafers into a solid resin negative mold, to increase their lifespan.

Fig7: Sublym – Alternative To Soft-Lithography
In 3D cell culture devices, so many variables have to be considered as we saw: origin, size, organization of the fibers… It can be interesting to have material like Flexdym, which is elastic, flexible, transparent. Besides, Flexdym does not require chemical treatment for the bounding to the class surface for instance.
Overall, microfluidics has greatly accelerated the advances in the field of 3D cell culture, we still need to move from the academic world to the patient. For this, it will be necessary to move from the laboratory to the industry and constantly improve knowledge in biology, engineering, materials …
References
[1] A. D. Castiaux, D.M. Spence, R.S. Martin, Anal. Methods, 2019, 11, 4220–4232. Read
[2] H. W. Wu, Y. H. Hsiao, C. C. Chen, S. F. Yet and C. H. Hsu, Molecules, 2016, 21, 1–11. Read
[3] F. F. Tao, X. Xiao, K. F. Lei and I. C. Lee, BioChip J., 2015, 9, 97–104. Read
[4] S. Chung, R. Sudo, P. J. Mack, C. R. Wan, V. Vickerman and R. D. Kamm, Lab Chip, 2009, 9, 269–275. Read
Dr. Constance Porrini
PharmD. and Ph.D. in Microbiology from National Research Institute for Agriculture, Food and the Environment (INRAE)

